This method is largely superseded by plot.PM_cov.
Usage
# S3 method for PMcov
plot(
x,
formula,
icen = "median",
include,
exclude,
mult = 1,
log = F,
square = F,
ref = F,
lowess = F,
grid = F,
ident = F,
reg = F,
ci = 0.95,
cex = 1,
cex.lab = 1.2,
x.stat = 0.6,
y.stat = 0.1,
col.stat = "black",
cex.stat = 0.8,
lwd = 2,
col = "red",
xlim,
ylim,
xlab,
ylab,
out = NA,
...
)Arguments
- x
The name of an PMcov data object generated by
makeCov- formula
This is a mandatory formula of the form
y ~ x, whereyandxare the twodataparameters to plot.- icen
A character vector to summarize covariate and parameter values. Default is “median”, but can also be one of “none”, “mean”. If
timeis a variable informula, the value will be set to “none” and theyvalues will be aggregated by subject ID vs. time.- include
A vector of subject IDs to include in the plot, e.g. c(1:3,5,15)
- exclude
A vector of subject IDs to exclude in the plot, e.g. c(4,6:14,16:20)
- mult
Multiplication factor for y axis, e.g. to convert mg/L to ng/mL
- log
Boolean operator to plot in log-log space; the default is
False- square
Boolean operator to force a square plot with equal x and y limits; the default is
True- ref
Boolean operator to draw a unity line; the default is
Trueunless “time” is the x value informulain which case this is ignored- lowess
Boolean operator to draw a lowess regression line; the default is
Falseand this is ignored if “time” is the x value informula- grid
Either a boolean operator to plot a reference grid, or a list with elements x and y, each of which is a vector specifying the native coordinates to plot grid lines; the default is
False. For example, grid=list(x=seq(0,24,2),y=1:10). Defaults for missing x or y will be calculated byaxTicks.- ident
Boolean operator to plot points as ID numbers; the default is
False. This option is useful to identify outliers.- reg
Boolean operator to draw a linear regression line; the default is
Trueunless “time” is the x value informulain which case this is ignored. If this option is selected, regression statistics will be printed on the plot if at least 3 subjects are included.- ci
The confidence interval for the linear regression parameter estimates; the default is 0.95.
- cex
Size of the plot symbols.
- cex.lab
Size of the plot labels.
- x.stat
Horizontal position to plot the linear regression statistics; the units are relative to the origin, i.e. extreme left is 0 and extreme right is 1.
- y.stat
Vertical position to plot the linear regression statistics; the units are relative to the origin, i.e. extreme bottom is 0 and extreme top is 1.
- col.stat
Color of the text for the regression statistics.
- cex.stat
Size of the text for the regression statistics.
- lwd
Width of the various regression or reference lines (unity, linear regression, or lowess regression)
- col
This parameter will be applied to the plotting symbol and is “red” by default.
- xlim
Limits of the x-axis as a vector, e.g.
c(0,1). It does not need to be specified, but can be.- ylim
Analogous to
xlim- xlab
Label for the x-axis. If missing, will default to the name of the x-variable.
- ylab
Label for the y-axis. If missing, will default to the name of the y-variable.
- out
Direct output to a PDF, EPS or image file. Format is a named list whose first argument,
typeis one of the following character vectors: “pdf”, “eps” (maps topostscript), “png”, “tiff”, “jpeg”, or “bmp”. Other named items in the list are the arguments to each graphic device. PDF and EPS are vector images acceptable to most journals in a very small file size, with scalable (i.e. infinite) resolution. The others are raster images which may be very large files at publication quality dots per inch (DPI), e.g. 800 or 1200. Default value isNAwhich means the output will go to the current graphic device (usually the monitor). For example, to output an eps file, out=list(“eps”) will generate a 7x7 inch (default) graphic.- ...
Other parameters as found in
plot.default.
Details
It will plot any two columns, specified using a formula, of a PMcov object,
which contains covariate and Bayesian posterior parameter information
for each subject. PMcov objects can be
accessed as the $data object within the $cov field of a PM_result object, e.g.
PM_result$cov$data.
Specifying any two variables that do not include time will result in a scatter plot with optional regression and reference lines. If
time is included as the x variable, the y variable will be plotted vs. time, aggregated by subject. This can be useful to see time varying parameters,
although a formula within formula approach may be required, e.g. plot(cov.1,I(cl_0*wt**0.75)~time) in order to see the change in cl over time according to
the change in wt over time, even though cl_0 is constant for a given subject.
Examples
library(PmetricsData)
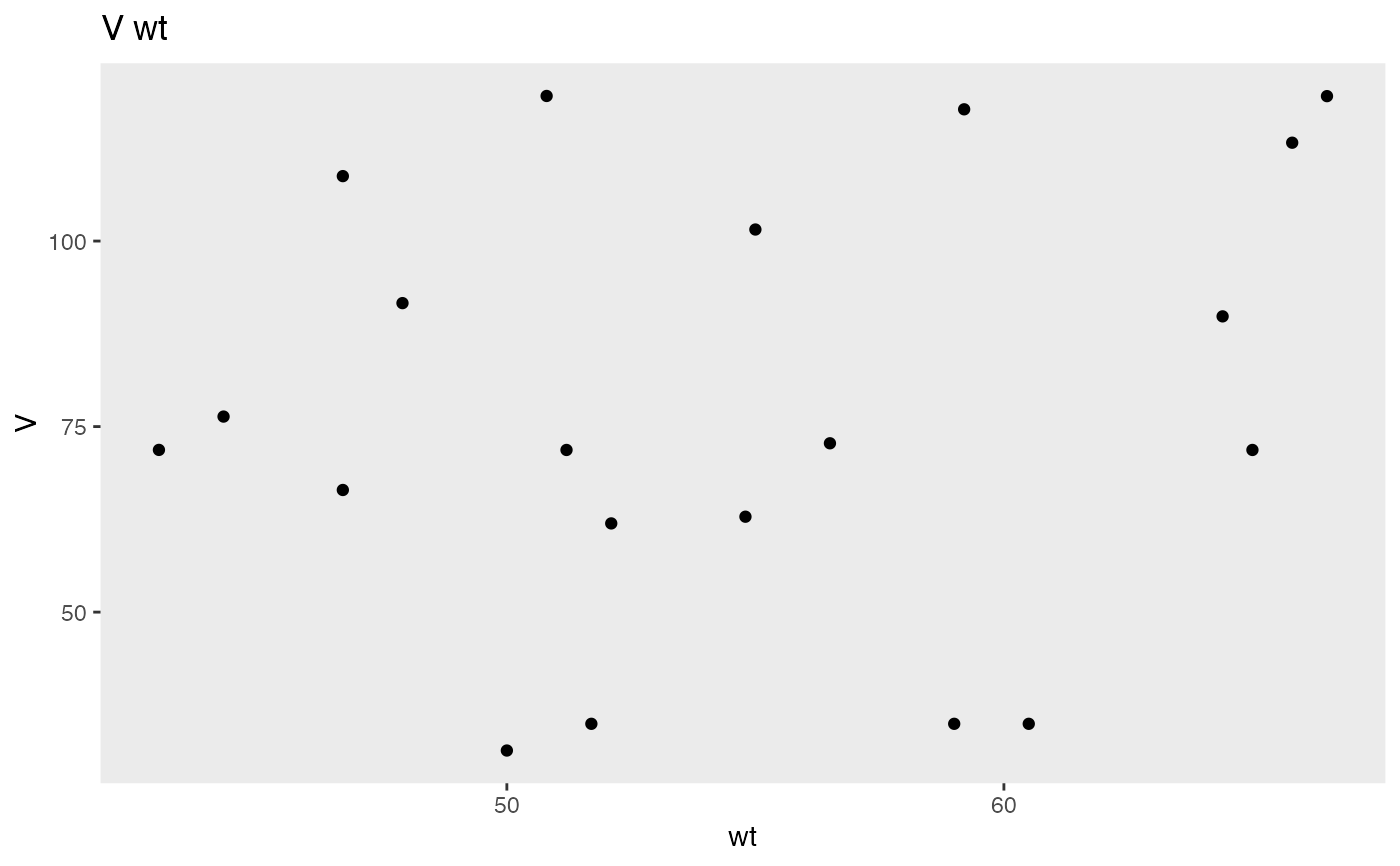
plot(NPex$cov$data, V ~ wt)